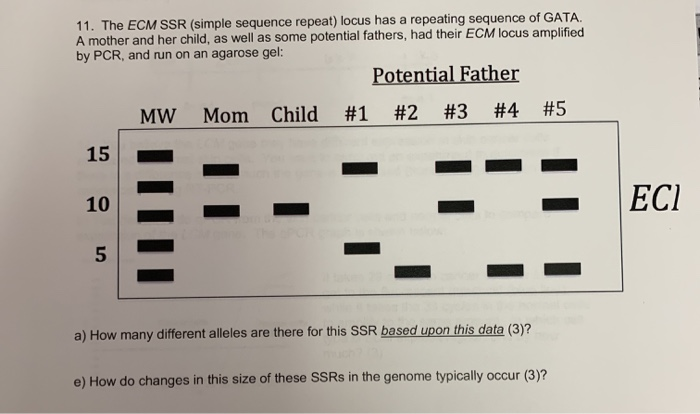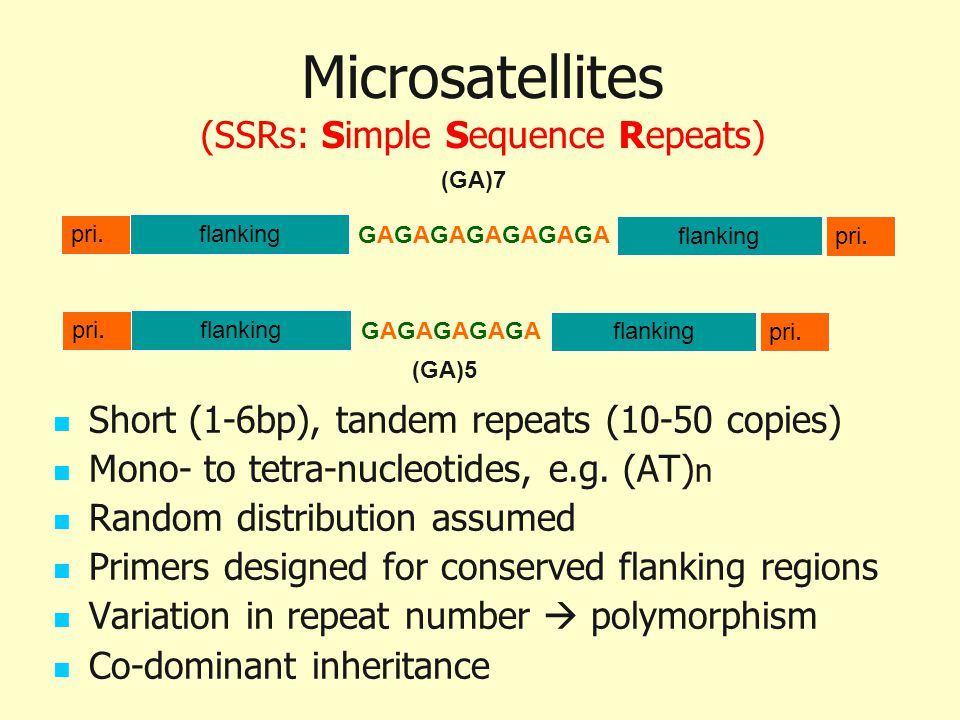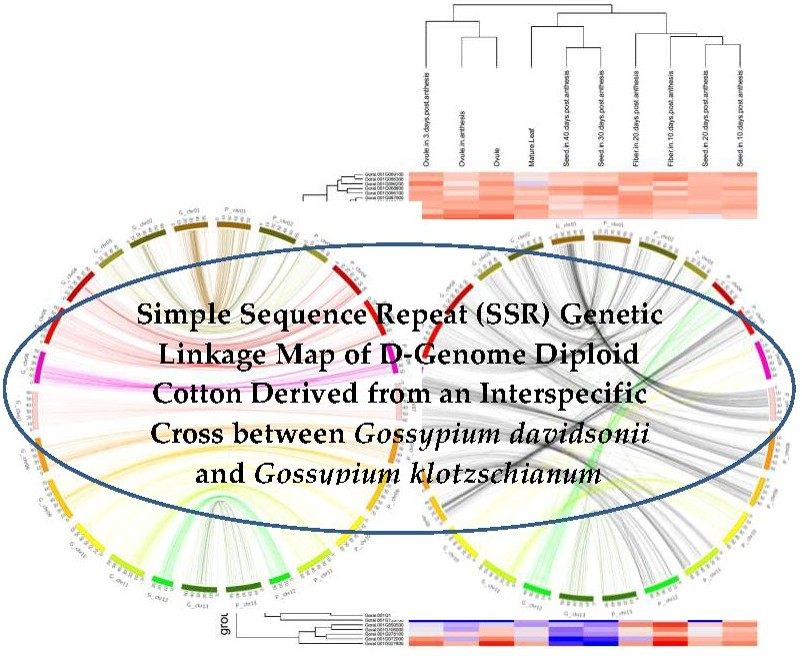
IJMS | Free Full-Text | Simple Sequence Repeat (SSR) Genetic Linkage Map of D Genome Diploid Cotton Derived from an Interspecific Cross between Gossypium davidsonii and Gossypium klotzschianum
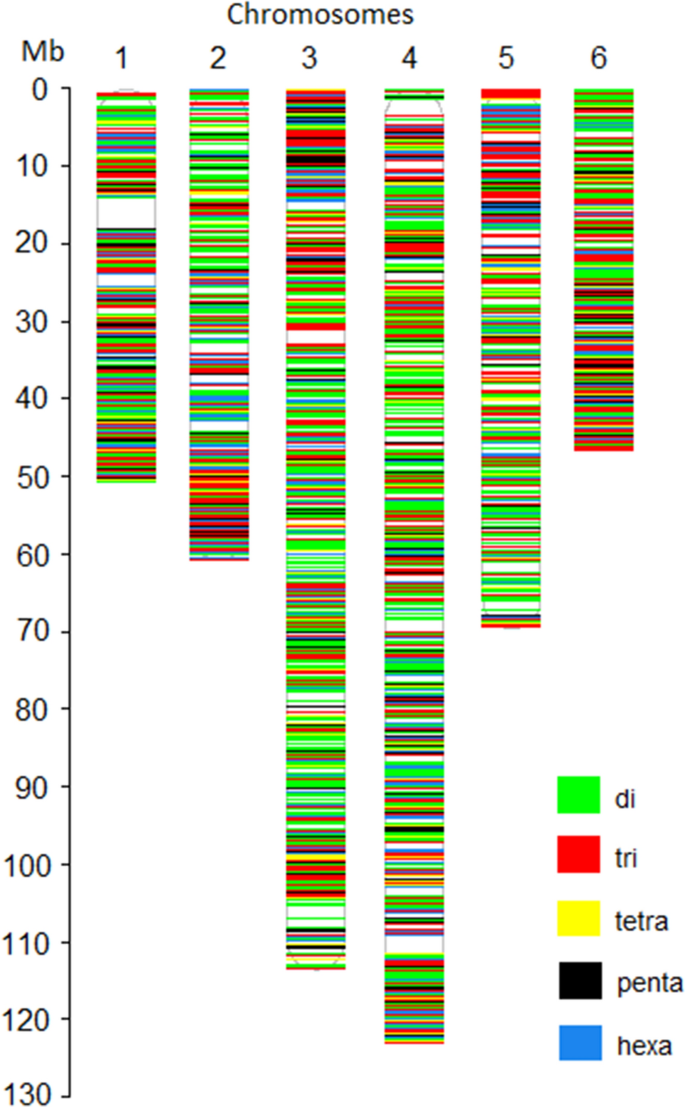
Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions | Scientific Reports

Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions | Scientific Reports
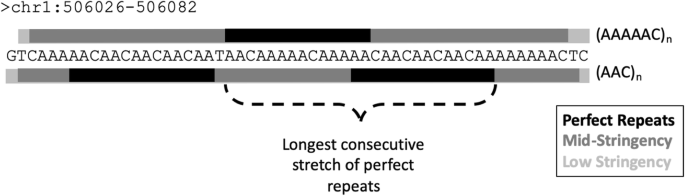
Finding and extending ancient simple sequence repeat-derived regions in the human genome | Mobile DNA | Full Text
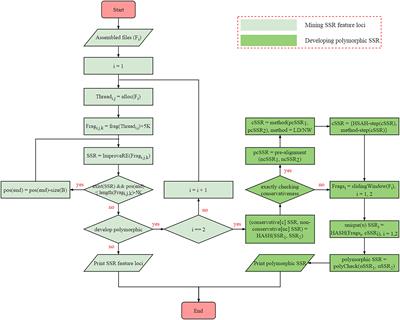
Frontiers | SSRMMD: A Rapid and Accurate Algorithm for Mining SSR Feature Loci and Candidate Polymorphic SSRs Based on Assembled Sequences
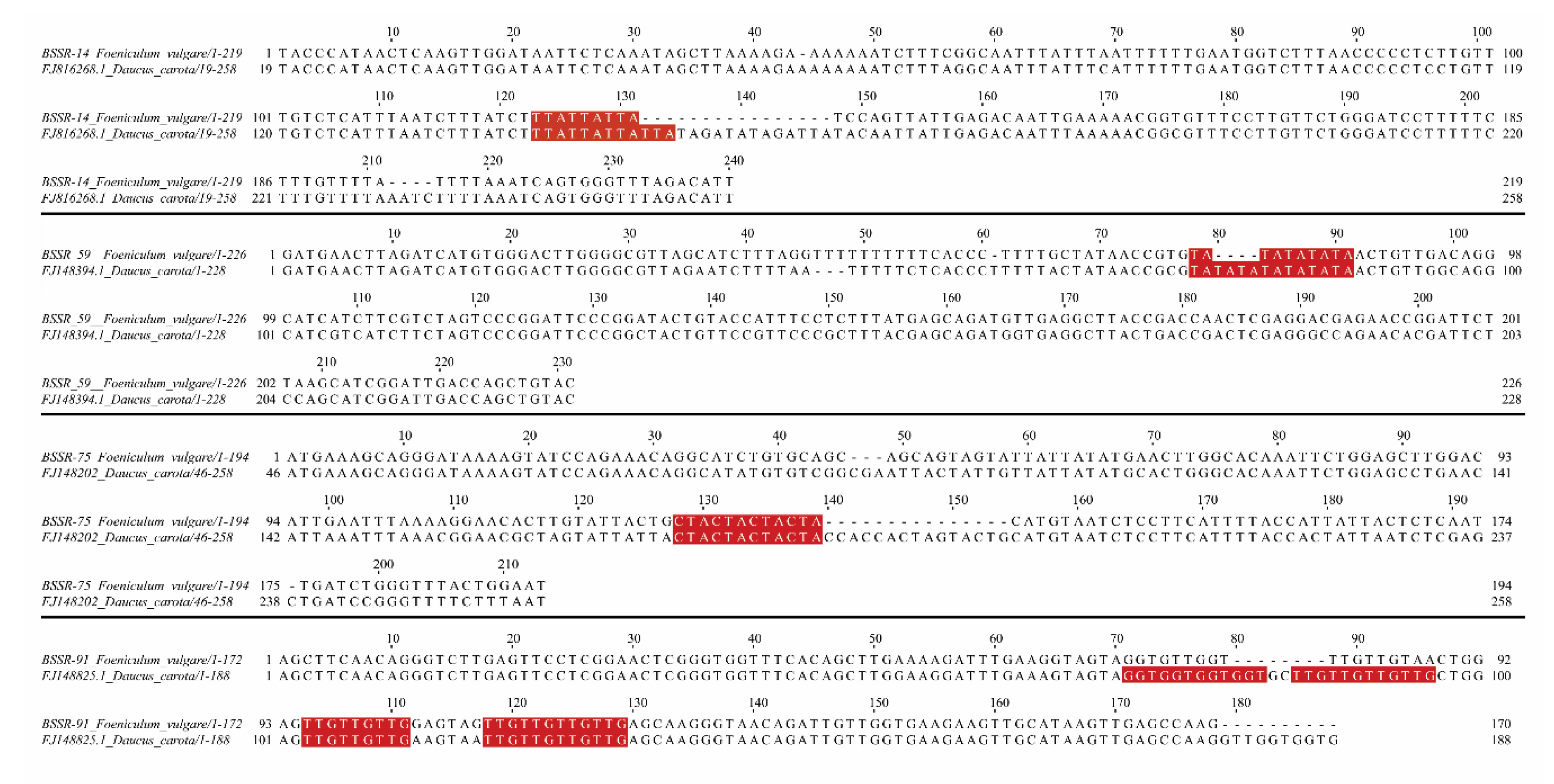

Plants | Free Full-Text | Evaluation of Cross-Species Transferability of SSR Markers in Foeniculum vulgare

Simple sequence repeat (SSR) analysis in the M. glyptostroboides cp... | Download Scientific Diagram
Fluorescent Inter Simple Sequence Repeat (F-ISSR) Markers and Capillary Electrophoresis to Assess Genetic Diversity and Relatedn
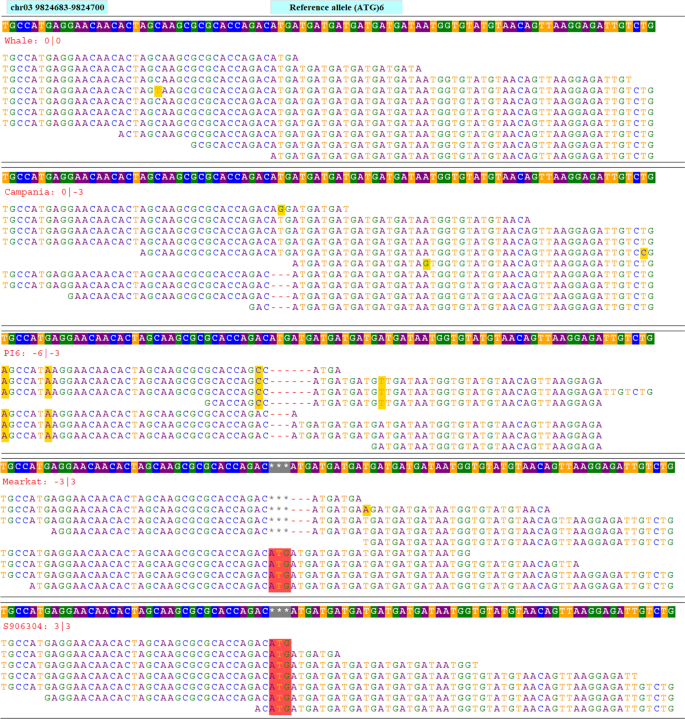
Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions | Scientific Reports

Figure 2 from Inter-Simple Sequence Repeat ( ISSR ) Markers : Are We Doing It Right ? | Semantic Scholar
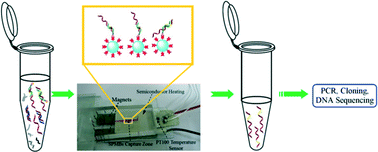
Fast magnetic isolation of simple sequence repeat markers in microfluidic channels - Lab on a Chip (RSC Publishing)

The Genotypes of Polymorphic Simple Sequence Repeat Loci Revealed by Whole-genome Resequencing Data of 30 Pyrus Accessions in: Journal of the American Society for Horticultural Science Volume 144 Issue 5 (2019)